Pyruvate kinase
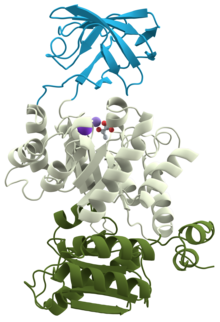
3D structure of pyruvate kinase
|
|||||||||
| Identifiers | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| EC number | 2.7.1.40 | ||||||||
| CAS number | 9001-59-6 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / EGO | ||||||||
|
|||||||||
| Search | |
|---|---|
| PMC | articles |
| PubMed | articles |
| NCBI | proteins |
Pyruvate kinase is the enzyme that catalyzes the final step of glycolysis. It catalyzes the transfer of a phosphate group from phosphoenolpyruvate (PEP) to adenosine diphosphate (ADP), yielding one molecule of pyruvate and one molecule of ATP. Pyruvate kinase is present in four distinct, tissue-specific isozymes, each consisting of particular kinetic properties necessary to accommodate the variations in metabolic requirements of diverse tissues.
There are four isozymes of pyruvate kinase in vertebrates: L (liver), R (erythrocytes), M1(muscles, hearts and brain) and M2 (only form detectable in early fetal tissue and present in most adult tissues). R and L isozymes differ from M1 and M2 in that they are both exclusively allosterically and reversibly regulated. From a kinetic standpoint, the R and L isozymes of pyruvate kinase have two key conformation states; one with a high substrate affinity and one with a low substrate affinity. The R-state, characterized by high substrate affinity, serves as the activated form of pyruvate kinase and is stabilized by PEP and FBP, promoting the glycolytic pathway. The T-state, characterized by low substrate affinity, serves as the inactivated form of pyruvate kinase, bound and stabilized by ATP and alanine, causing phosphorylation of pyruvate kinase and the inhibition of glycolysis.
Gene expression varies between the different isozymes. M1 and M2 isozymes are regulated by the gene PKM and R and L isozymes are regulated by the gene PKLR. In terms of structure, there is both a tetrameric and dimeric form of pyruvate kinase. The tetrameric form is the pyruvate kinase structure in its R-state conformation, namely with high binding affinity to PEP. In contrast, the dimeric form is its structure in T-state conformation, namely with a low binding affinity to PEP. As a result, gene expression can be regulated by converting the highly active tetrameric form of PKM2, which yields high PEP concentrations, into an inactive dimeric form, which yields a PEP concentration of nearly zero.
Many Enterobacteriaceae, including E. coli, have two isoforms of pyruvate kinase, PykA and PykF, which are 37% identical in E. coli (Uniprot: PykA, PykF). They catalyze the same reaction as in eukaryotes, namely the generation of ATP from ADP and PEP, the last step in glycolysis, a step that is irreversible under physiological conditions. PykF is allosterically regulated by fructose 1,6-bisphosphate (FBP) which reflects the central position of PykF in cellular metabolism. PykF transcription in E. coli is regulated by the global transcriptional regulator, Cra (FruR). PfkB was shown to be inhibited by MgATP at low concentrations of Fru-6P, and this regulation is important for gluconeogenesis.
...
Wikipedia
