Aspartate protease
| Eukaryotic aspartyl protease | |||||||||
|---|---|---|---|---|---|---|---|---|---|

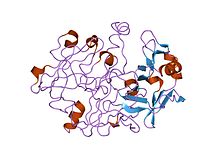
Structure of the dimeric aspartic protease HIV protease in white and grey, with peptide substrate in black and active site aspartate side chains in red. (PDB: 1KJF)
|
|||||||||
| Identifiers | |||||||||
| Symbol | Asp | ||||||||
| Pfam | PF00026 | ||||||||
| InterPro | IPR001461 | ||||||||
| PROSITE | PDOC00128 | ||||||||
| SCOP | 1mpp | ||||||||
| SUPERFAMILY | 1mpp | ||||||||
| OPM superfamily | 108 | ||||||||
| OPM protein | 1lyb | ||||||||
|
|||||||||
| Available protein structures: | |
|---|---|
| Pfam | structures |
| PDB | RCSB PDB; PDBe; PDBj |
| PDBsum | structure summary |
| A1_Propeptide | |||||||||
|---|---|---|---|---|---|---|---|---|---|
crystal and molecular structures of human progastricsin at 1.62 angstroms resolution
|
|||||||||
| Identifiers | |||||||||
| Symbol | A1_Propeptide | ||||||||
| Pfam | PF07966 | ||||||||
| InterPro | IPR012848 | ||||||||
|
|||||||||
| Available protein structures: | |
|---|---|
| Pfam | structures |
| PDB | RCSB PDB; PDBe; PDBj |
| PDBsum | structure summary |
Aspartic proteases are a catalytic type of protease enzymes that use an activated water molecule bound to one or more aspartate residues for catalysis of their peptide substrates. In general, they have two highly conserved aspartates in the active site and are optimally active at acidic pH. Nearly all known aspartyl proteases are inhibited by pepstatin.
Aspartic endopeptidases EC 3.4.23. of vertebrate, fungal and retroviral origin have been characterised. More recently, aspartic endopeptidases associated with the processing of bacterial type 4 prepilin and archaean preflagellin have been described.
Eukaryotic aspartic proteases include pepsins, cathepsins, and renins. They have a two-domain structure, arising from ancestral duplication. Retroviral and retrotransposon proteases (retroviral aspartyl proteases) are much smaller and appear to be homologous to a single domain of the eukaryotic aspartyl proteases. Each domain contributes a catalytic Asp residue, with an extended active site cleft localized between the two lobes of the molecule. One lobe has probably evolved from the other through a gene duplication event in the distant past. In modern-day enzymes, although the three-dimensional structures are very similar, the amino acid sequences are more divergent, except for the catalytic site motif, which is very conserved. The presence and position of disulfide bridges are other conserved features of aspartic peptidases.
Aspartyl proteases are a highly specific family of proteases – they tend to cleave dipeptide bonds that have hydrophobic residues as well as a beta-methylene group. Unlike serine or cysteine proteases these proteases do not form a covalent intermediate during cleavage. Proteolysis therefore occurs in a single step.
...
Wikipedia
