Leucine rich repeat
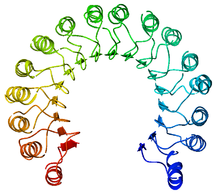
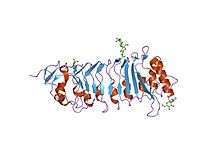
A leucine-rich repeat (LRR) is a protein structural motif that forms an α/β horseshoe fold. It is composed of repeating 20–30 amino acid stretches that are unusually rich in the hydrophobic amino acid leucine. These repeats commonly fold together to form a solenoid protein domain, termed leucine-rich repeat domain. Typically, each repeat unit has beta strand-turn-alpha helix structure, and the assembled domain, composed of many such repeats, has a horseshoe shape with an interior parallel beta sheet and an exterior array of helices. One face of the beta sheet and one side of the helix array are exposed to solvent and are therefore dominated by hydrophilic residues. The region between the helices and sheets is the protein's hydrophobic core and is tightly sterically packed with leucine residues.
Leucine-rich repeats are frequently involved in the formation of protein–protein interactions.
Leucine-rich repeat motifs have been identified in a large number of functionally unrelated proteins. The best-known example is the ribonuclease inhibitor, but other proteins such as the tropomyosin regulator tropomodulin and the toll-like receptor also share the motif. In fact, the toll-like receptor possesses 10 successive LRR motifs which serve to bind pathogen- and danger-associated molecular patterns.
Although the canonical LRR protein contains approximately one helix for every beta strand, variants that form beta-alpha superhelix folds sometimes have long loops rather than helices linking successive beta strands.
...
Wikipedia